Transcriptomic analysis
The importance of gene expression in the study of different diseases.
Home » Transcriptomics
Study of gene expression
Transcriptomic analyses allow quantification of gene expression using NGS platforms. The interpretation of data from RNA sequencing (RNA-Seq) has become an increasingly demanded service due to its sensitivity and accuracy.
Our bioinformatics analysis includes:
- Quality control of the sequences.
- Alignment of reads against reference sequences.
- Quantification of gene expression in the sample.
- Analysis of differential gene expression between different samples.
- Enrichment study of gene ontologies and pathways.
- Study of isoforms generated in alternative splicing events*.
- Identification of other RNAs (smallRNAs and ncRNAs)*.
*Where the experimental design and coverage allow.
Documentation
We can carry out bioinformatics analysis from:
- Illumina: FASTQ
- MGI: FASTQ
You can send us your raw files on a hard disk or share them via FTP. If you have any questions please contact us by sending an email to info@dreamgenics.com or calling 985 088 180 and we will help you.
Results in Genome One Reports
We deliver the results of our transcriptomic analysis through our Genome One Reports platform. A simple and intuitive web platform that will provide you with the following advantages:
- You will have access to the quality analysis of the sequences obtained.
- Sample correlation analysis and differential expression study.
- PCA plot, Adjusted p-value histogram, MA plot, Volcano plot, Heatmap and Correlation heatmap
- Enrichment of Gene Ontologies paths and terms (WikiPathways, GO-MF, GO-CC and GO-BP)
- All tables and figures are easily exportable, ready to be used in scientific publications.
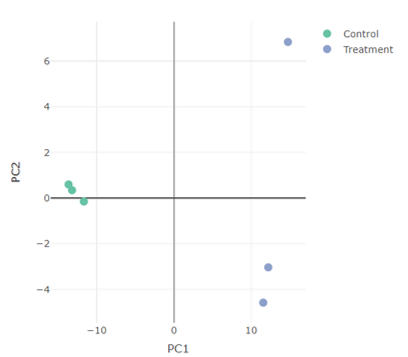
PCA Plot
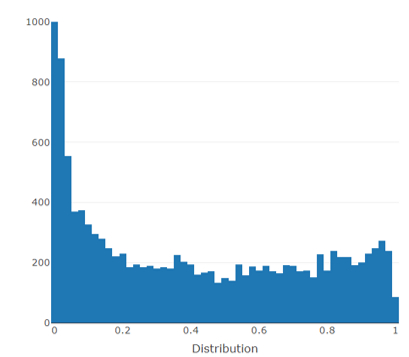
Adjusted P-value Histogram
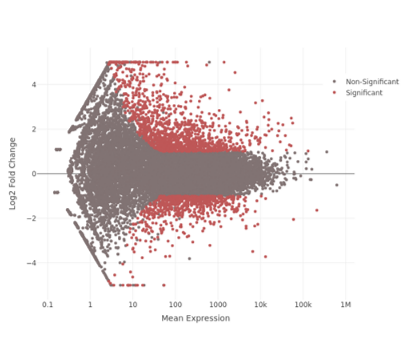
MA Plot
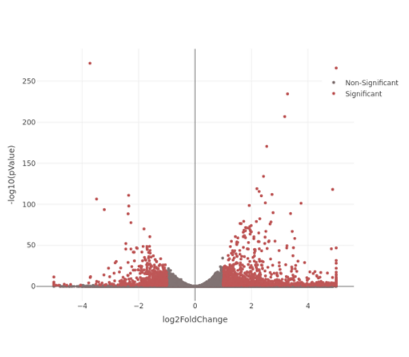
Volcano Plot
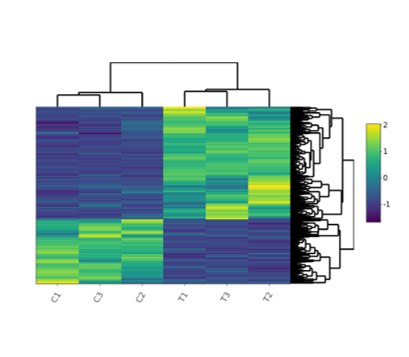
Heatmap
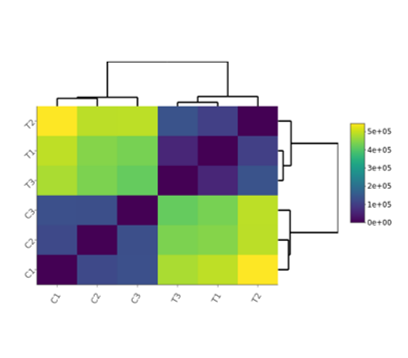
Correlation Heatmap
Different options in RNA-Seq experiments
The libraries used in an RNA-Seq experiment depend on the starting sample and the type of RNAs to be studied.
Contact us and we will help you choose the best option for your project.
mRNA-Seq
- NovaSeq 6000 Platform
- TruSeq Stranded mRNA library
- mRNA analysis
- >1ug of RNA with RIN>7
- 30-40 M/sample
- Minimum 3 samples per condition
- Differential expression, ontologies and pathways
- Results in Genome One Reports
Total RNA-Seq
- NovaSeq 6000 Platform
- TruSeq Stranded Total RNA Library
- mRNA and lncRNA analysis
- >1ug of RNA with RIN>7
- 40-60 M/sample
- Ribo-Zero Globin* treatment
- Minimum 3 samples per condition
- Differential expression, ontologies and pathways
- Results in Genome One Reports
*Elimination of ribosomal RNA and globin mRNA in blood samples.




















